Guiding Intracardiac Beating Heart Procedures With Real-Time Three-Dimensional Ultrasound
Paul NovotnyMotivation
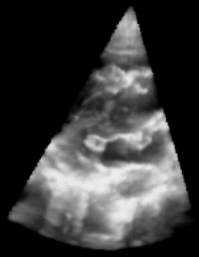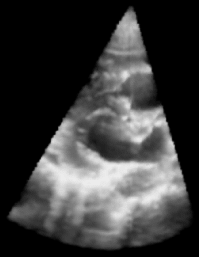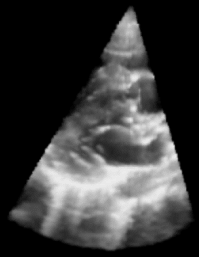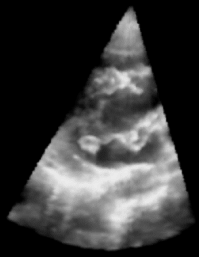 |
| Fig. 1: 3D Ultrasound image of a beating heart. |
3D Ultrasound
Traditional 3D ultrasound systems rely on slow offline image processing to construct a 3D volume from a series of 2D ultrasound images. Although, these systems produce good three-dimensional images, they are far from real time, often requiring minutes to construct. The time lag prohibits use in interventional procedures where accurate up to date information is critical for task performance. Phillips Medical Systems has recently released a system that produces 3D ultrasound images in real time (20-25 frames per second). The system consists of a fully sampled two-dimensional array of ultrasound transducers that quickly scans a three dimensional volume. These characteristics suggest that the Philips 3D ultrasound system is a feasible imaging modality for guiding surgical tasks.
Previous Work
In earlier work, our group demonstrated that 3D ultrasound could guide complex surgical tasks in vitro[1]. This was validated by seven surgical trainees performing a range of tasks in a testing tank. The subjects performed these tasks using 2D ultrasound, 3D ultrasound, or endoscopic video for guidance.
 |
| Fig. 2: Picture of clip fixation task with corresponding 3D ultrasound image. |
- Bead in Hole
Using an endoscopic grasper, the subject tries to place a 4mm bead into a 5mm hole cut out of a canvas sheet. - Point to Point Navigation
In this task, the surgeons were asked to navigate the tip of a blunt dissection tool from one point in the tank to another.
- Clip Fixation
The goal of this task is to grasp two surgical tubes with an endoscopic tool and then apply a metal clip with an endoscopic clipping tool. This task is a simulation of a common surgical maneuver to attach adjacent tissues.
Ultrasound and Surgical Instruments
It is becoming increasingly clear, that instruments
such as endoscopic graspers when immersed in liquids display artifacts and irregularities
that make it difficult to determine the tool’s location, orientation, and geometry.
In addition, the instrument’s shape can appear incomplete due to its orientation or
obstacles in the field. Since the geometry and properties of these instruments are
known a priori, it is feasible to combine this information with the ultrasound image
data to locate and render an enhanced representation of the instrument.
 |
Fast block flow tracking of atrial septal defects in 4D echocardiography
Marius George LinguraruWe have developed beating-heart atrial septal defect (ASD) closure techniques using real-time 3D ultrasound guidance. The major image processing challenges are the low-image quality and the processing of information at high-frame rate. We introduce a block flow technique, which combines the velocity computation from optical flow for an entire block with template matching. Enforcing adapted similarity constraints to both the previous and first frames ensures optimal and unique solutions. We compare the performance of the proposed algorithm with that of block matching and region-based optical flow on eight in vivo 4D datasets acquired from porcine beating-heart procedures. Results show that our technique is more stable and has higher sensitivity than both optical flow and block matching in tracking ASDs. Computing velocity at the block level, our technique tracks ASD motion at 2 frames/s, much faster than optical flow and comparable in computation cost to block matching, and shows promise for real-time (30 frames/s). We report consistent results on clinical intra-operative images and retrieve the cardiac cycle (in ungated images) from error analysis. Quantitative results are evaluated on synthetic data with maximum tracking errors of 1 voxel.
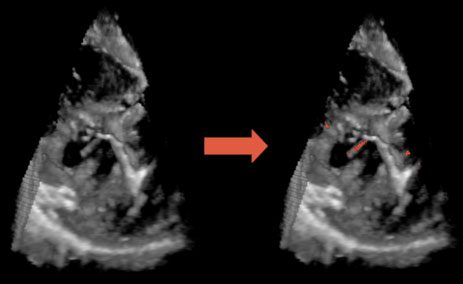
| Fig. 1. 3D US image of a porcine beating-heart with ASD. The US probe is placed outside the right atrium and the beam oriented toward the left atrium. (a) shows the entire 3D US volume, while (b) presents a magnified view of the ASD. The patch must cover the entire ASD surface, as seen in (c). |
| Fig. 2. The comparative performance of the four tracking algorithms. The top row shows results by block matching (BM); the second row by optical flow (OF); the third row by a multi-scale combination of block-matching and optical flow (BMOF); the bottom row by block flow (BF). All blocks are shown equally sized. From left to right, the columns show 3D blocks at frames 1, 20, 40 and 50. |
A Nonrigid Image Registration Framework for Identification of Tissue Mechanical Parameters
Petr JordanWe present a modular framework for mechanically regular ized nonrigid image registration of 3D ultrasound and for identification of tissue mechanical parameters. Mechanically regularized deformation fields are computed from sparsely estimated local displacements. We enforce image-based local motion estimates by applying concentrated forces at mesh nodes of a mechanical finite-element model. The concentrated forces are generated by the elongation of regularization springs connected to the mesh nodes as their free ends are displaced according to local motion estimates. The regularization energy corresponding to the potential energy stored in the springs is minimized when the mechanical response of the model matches the observed response of the organ. We demonstrate that this technique is suitable for identification of material parameters of a nonlinear viscoelastic liver model and demonstrate its benefits over traditional indentation methods in terms of improved volumetric agreement between the model response and the experiment.
| Fig. 1. In the proposed mechanical regularization framework, sparse local motion estimates are coupled to a mechanical finite-element model as lumped body forces applied by displaced regularization springs. This results in a mechanically constrained deformation field uFEM and regularization energy. |
| Fig. 2. Left: A cut through the deforming finite-element model and 3DUS sequence. Middle: Slice through the undeformed 3DUS volume. Right: Slice through the deforming volume (t=1.2 s), showing the local optical flow and the 3 locations evaluated. |